Ask Questions, Get Insights
Conversational Lung
Biology AI
Perform complex multiomics analyses through natural language. Integrate single-cell RNA-seq, spatial transcriptomics, pathway enrichment, drug repurposing, and literature research—no coding required. Accelerate discovery with publication-ready results.
About LungChat
Democratizing access to complex lung biology analysis
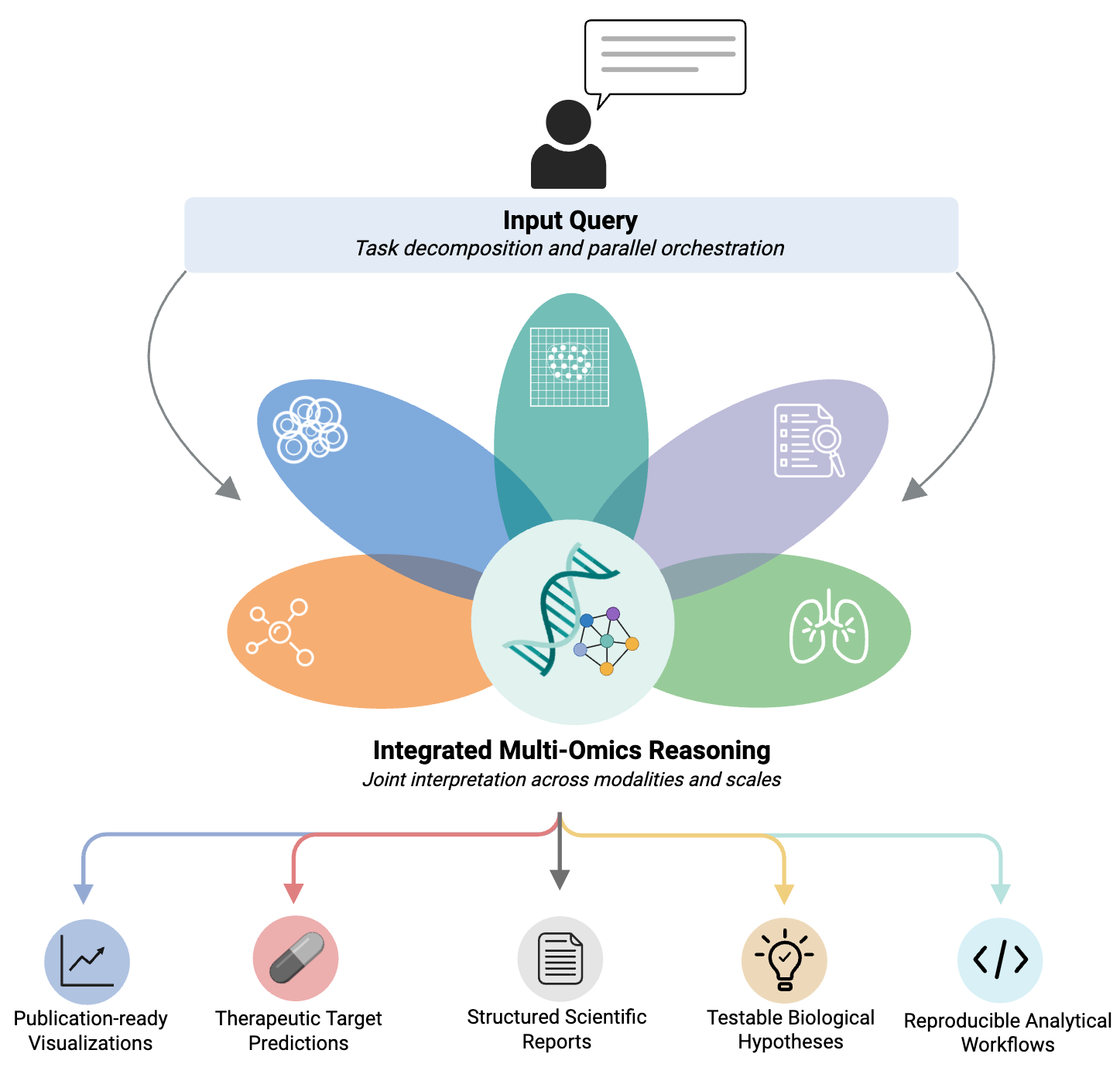
LungChat democratizes access to advanced multiomics analysis for lung biology research. Simply ask questions in natural language—LungChat automatically decomposes complex queries into subtasks and orchestrates parallel analysis across multiple specialized agents. Seamlessly integrate single-cell RNA-seq data from 470+ donors, spatial transcriptomics, pathway enrichment from 120+ databases, drug connectivity analysis from 143K+ compounds, and literature synthesis—all without writing a single line of code.
Through integrated multi-omics reasoning, LungChat jointly interprets findings across modalities and scales—connecting cellular signatures to spatial niches, pathways to therapeutic targets, and discoveries to published literature. The result: actionable scientific insights with full reproducibility.
Scientific Capabilities
Unleashing new possibilities for lung biology research
Comprehensive Single-Cell Analysis
Query 2.8M+ cells from 470+ donors across 40+ studies covering normal lung, development, and 15+ diseases (IPF, COPD, COVID-19, BPD, lung cancer). Generate publication-ready UMAPs, heatmaps, violin plots, volcano plots, dotplots, and differential expression analyses. Perform cross-dataset comparisons to identify conserved vs. disease-specific signatures, with statistical cell type proportion testing and interactive web-based heatmap exploration.
Spatial Transcriptomics & Cell Communication
Analyze spatial data from Xenium and Visium platforms to map cellular niches and tissue architecture. Explore cell-cell communication with CellChat—visualize ligand-receptor interactions as chord diagrams and signaling pathway networks. Perform spatial neighborhood enrichment with Squidpy, cross-platform Xenium-Visium correlation analysis, and integrate H&E/DAPI histology for multi-modal figures.
Drug Discovery & Repurposing
Identify potential therapeutics using connectivity mapping against 143K+ drug perturbation profiles. Find compounds that could reverse disease gene signatures, investigate drug mechanisms of action, explore gene knockout effects, and discover structurally similar compounds. Analyze drug impact across specific cell types for precision medicine applications.
Functional Enrichment & Literature
Perform pathway enrichment using 120+ databases (GO, KEGG, Reactome, disease ontologies, drug targets) to identify biological processes, molecular functions, and disease associations. Search PubMed with intelligent relevance ranking that prioritizes clinical trials and experimental evidence. Synthesize findings to contextualize discoveries and connect pathways to disease mechanisms.
Advanced Visualization Suite
Create gene-gene interaction networks, correlation matrices, and Venn/UpSet diagrams for gene set comparisons. Generate radar charts for cell composition, box plots with statistical testing for cell frequency analysis, and multi-panel spatial figures combining Xenium, Visium, H&E and DAPI. All visualizations are publication-ready in PNG and PDF formats.
LungMAP Database Integration
Query the LungMAP consortium database through natural language to access comprehensive lung development and disease resources. Discover datasets, explore sample demographics, retrieve gene lists from computational analyses, and perform cross-species comparisons for developmental trajectory analysis.
Custom Computational Analysis
Execute custom Python or R code in a secure sandbox for novel analyses, statistical computations, data transformations, and custom visualizations. Process TSV/CSV files, perform statistical tests, and create publication-quality plots tailored to your specific research questions.
Reproducibility & Provenance
Every analysis generates publication-ready figures (PNG/PDF), data tables (TSV), and complete parameter logs (JSON) ensuring full reproducibility and scientific rigor. All outputs are downloadable for publication, sharing, and exact replication of analyses.
Our Team
Meet the researchers behind LungChat
Frequently Asked Questions
Find answers to common questions about LungChat
LungChat integrates single-cell RNA-seq, spatial transcriptomics, drug repurposing, cell-cell communication analysis, functional enrichment, and literature research in a unified natural language interface. Unlike traditional tools that require coding expertise, LungChat enables researchers to perform complex multiomics analyses through conversational queries—from differential expression to finding therapeutic candidates—while ensuring full reproducibility with complete parameter tracking and publication-ready outputs.
LungChat integrates 2.8M+ cells from 470+ donors across 40+ single-cell RNA-seq studies covering normal lung, development, and 15+ disease states (IPF, COPD, COVID-19, BPD, lung cancer, and more). Datasets include the Human Lung Cell Atlas (HLCA) with 35 author studies, fetal development atlases, and disease-specific cohorts. Spatial transcriptomics data includes Xenium and Visium platforms with multi-resolution annotations for tissue niches, cellular communication analysis with CellChat, and integrated histology imaging.
Yes, LungChat is freely available for research purposes. To ensure fair access and system stability, each user can submit up to 15 queries per day and 50 queries per week.
Every analysis generates publication-ready figures (PNG/PDF), data tables (TSV), and complete parameter logs (JSON) containing all analysis parameters, dataset identifiers, and tool configurations. This ensures full provenance tracking and allows anyone to reproduce the exact analysis, meeting the highest standards of scientific reproducibility.
LungChat uses connectivity mapping to match your disease gene signatures against 143K+ drug perturbation profiles from the iLINCS database. It identifies compounds whose effects could reverse disease signatures, helping prioritize candidates for therapeutic development. You can also investigate specific drug mechanisms, explore gene knockout effects, and discover structurally similar compounds—all through natural language queries.
Currently, LungChat works with pre-integrated datasets from the LungMAP consortium and published studies. Support for user-uploaded datasets is planned for future releases. Contact us if you have specific datasets you'd like integrated or if you're interested in collaborating.
Research Paper
Our preprint describing LungChat's capabilities and scientific applications is coming soon.
Coming SoonGet in Touch
Have questions? We're here to help

